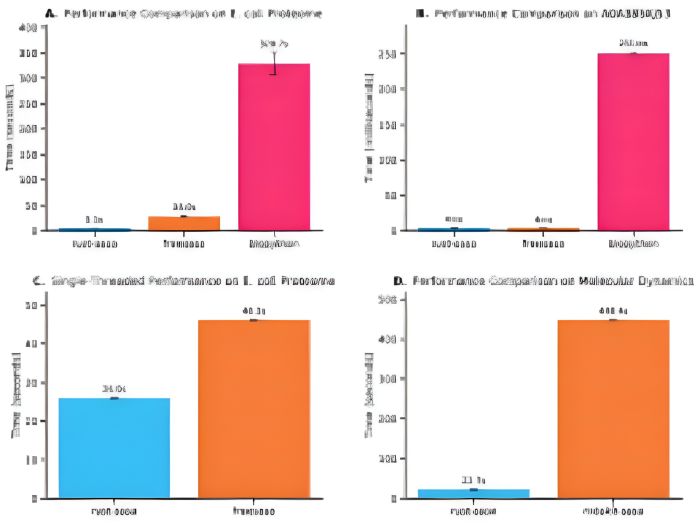

Solvent accessible surface area (SASA) calculations are fundamental for understanding protein structure, function, and dynamics in computational biology, providing insights into protein folding, stability, and intermolecular interactions. The Shrake-Rupley algorithm has served as the standard method since 1973, but existing implementations often become computational
bottlenecks when analyzing large protein datasets or molecular dynamics trajectories.
Something went wrong, please try again later.
This resource hasn't been reviewed yet
To ensure quality for our reviews, only customers who have purchased this resource can review it
Report this resourceto let us know if it violates our terms and conditions.
Our customer service team will review your report and will be in touch.